Transcription
Transcription = synthesis of RNA using a DNA template.
RNA polymerase binds to DNA, separates the strands locally, and builds an RNA strand by joining complementary RNA nucleotides.
Base-pairing rule in transcription: A on DNA template pairs with U on RNA, C pairs with G.
The DNA template strand is read, but the DNA base sequence itself is conserved: DNA is stable and can be reused many times.
This stability is essential because cells, especially long-lived somatic cells, must keep important DNA sequences unchanged.
Not all genes are expressed all the time; transcription is a key control point in gene expression.
Typical exam trap: do not say transcription makes protein directly; it makes RNA, usually mRNA.
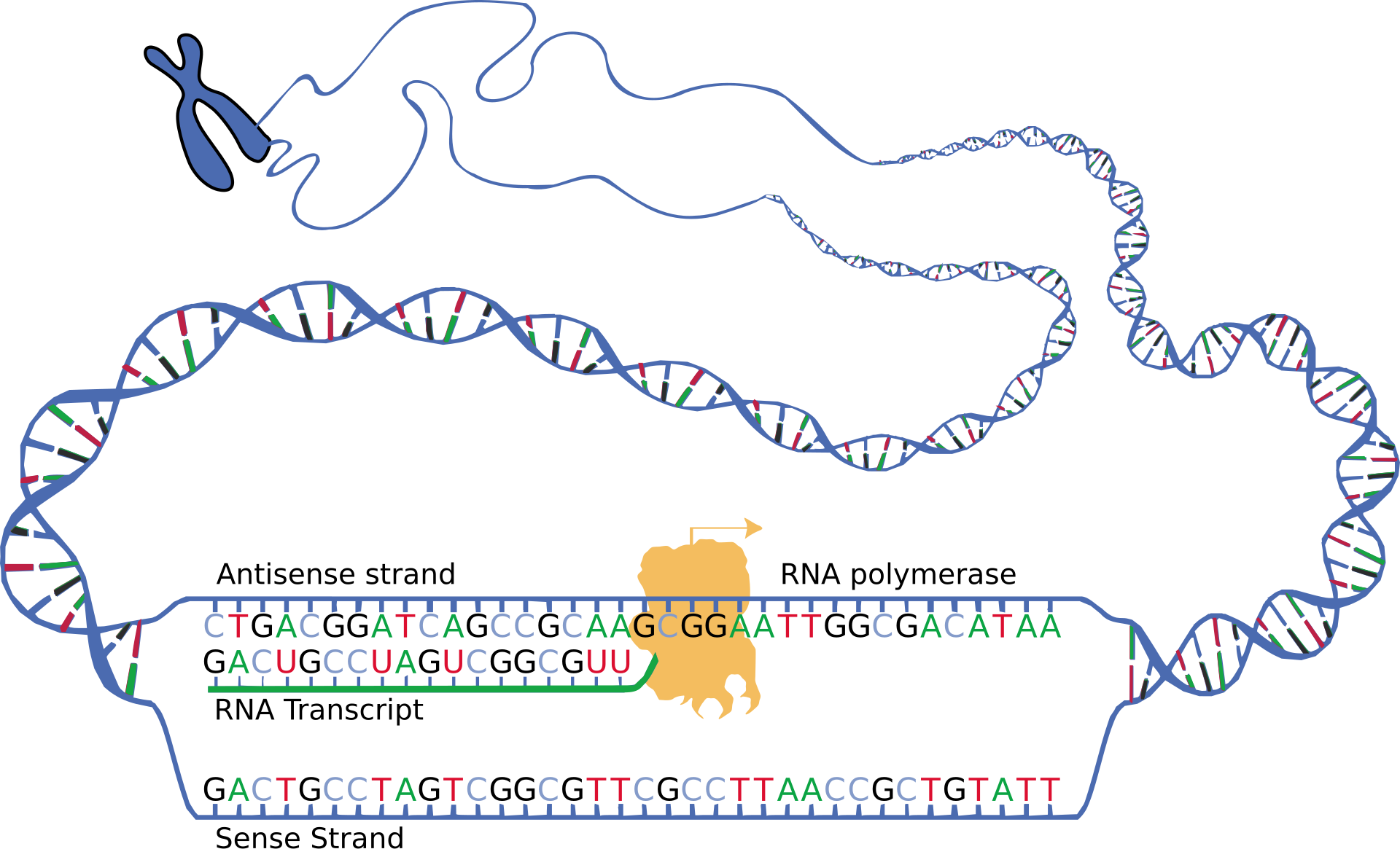
This diagram shows RNA polymerase using one DNA strand as a template to build a complementary RNA transcript. It helps visualize why U pairs with A in RNA and why transcription copies information without changing the DNA template. Source
Translation
Translation = synthesis of a polypeptide from an mRNA sequence.
The mRNA binds to the small subunit of the ribosome.
The large ribosomal subunit allows two tRNAs to bind at the same time during elongation.
tRNA carries specific amino acids and has an anticodon that pairs with an mRNA codon.
Codon = base triplet on mRNA; anticodon = complementary base triplet on tRNA.
Correct codon–anticodon complementary base pairing ensures the correct amino acid is added.
The ribosome moves stepwise along the mRNA, reading codons in sequence.
Amino acids are linked by peptide bonds to form the growing polypeptide chain.
IB emphasis is on elongation: ribosome movement, tRNA pairing, and peptide bond formation.
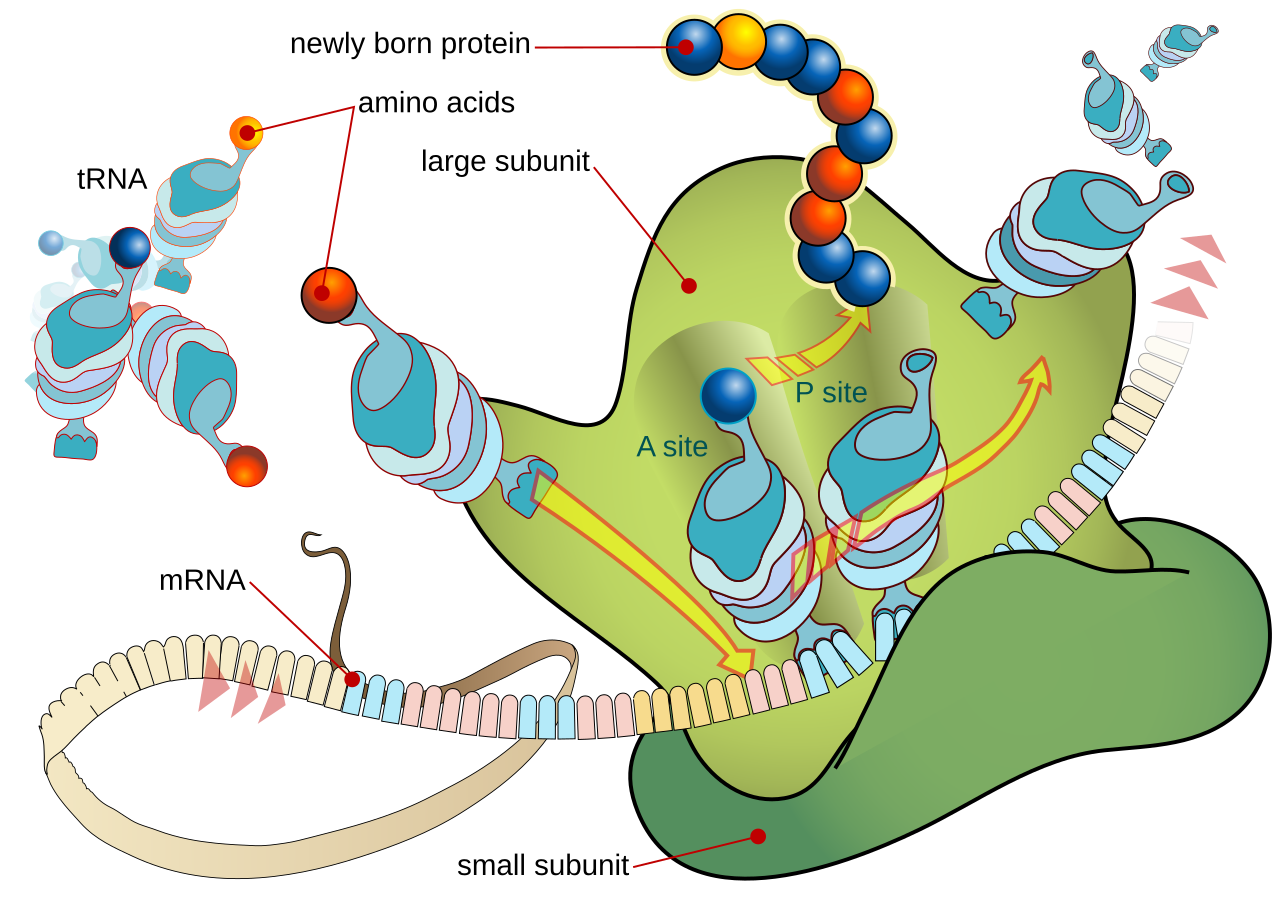
This image shows a ribosome, mRNA, and multiple tRNA molecules interacting during translation. It is useful for explaining codon–anticodon pairing and how amino acids are assembled in the correct order. Source
The genetic code
The genetic code is read in triplets called codons.
A triplet code is needed because there are 4 bases but 20 amino acids, so single bases or pairs would not provide enough combinations.
The code is degenerate: more than one codon can code for the same amino acid.
The code is universal: almost all organisms use the same codon meanings, supporting common ancestry.
Students must be able to use a table of mRNA codons to deduce the amino acid sequence.
Always read a codon table using mRNA codons, not DNA triplets, unless the question asks you to convert first.
Common exam skill: convert DNA template → mRNA → amino acid sequence.
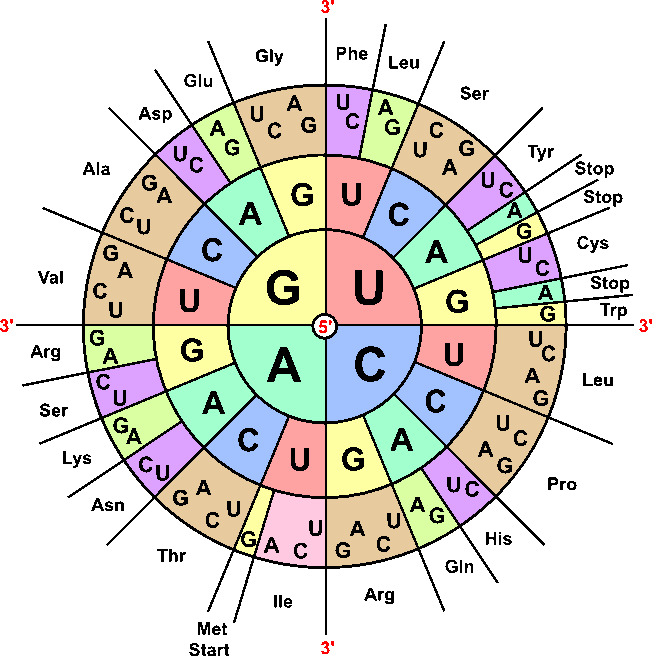
This codon table shows which mRNA codons specify each amino acid and where stop codons occur. It is ideal for practice in translating an mRNA sequence into a polypeptide sequence in exam questions. Source
How sequence determines protein structure
The base sequence of a gene determines the base sequence of mRNA.
The mRNA codon sequence determines the amino acid sequence of the polypeptide.
Therefore, a change in DNA can cause a change in primary structure of a protein.
A different amino acid sequence can alter folding and therefore protein structure and function.
Mutations that change protein structure can therefore affect phenotype.
A useful example is a point mutation that changes one codon, causing a different amino acid to be inserted.
Because the code is degenerate, some base substitutions may be silent and not change the amino acid.
Mutations and protein synthesis
A mutation can alter the mRNA codon sequence produced during transcription.
During translation, this may change the amino acid sequence and therefore the final protein.
Point mutations are especially important in exam questions because students may need to compare normal and mutant sequences.
Mutation effects may include:
No change to amino acid sequence (silent mutation) because of degeneracy.
One amino acid changed, potentially altering protein function.
Premature stop codon, producing a shorter polypeptide.
Always follow the sequence logic step by step: DNA → mRNA → codons → amino acids → likely protein effect.
HL only: directionality and transcription control
Transcription and translation both proceed in a 5' to 3' direction.
In transcription, RNA nucleotides are added to the 3' end of the growing RNA strand, so the RNA is synthesized 5' → 3'.
In translation, the ribosome moves along mRNA in the 5' → 3' direction.
Transcription begins at a promoter.
Transcription factors can bind to the promoter and help initiate transcription.
Important exam distinction: many DNA sequences are non-coding and do not code for polypeptides.
Examples of non-coding DNA to know: regulators of gene expression, introns, telomeres, and genes for rRNA and tRNA in eukaryotes.
HL only: RNA processing in eukaryotes
In eukaryotes, the initial RNA transcript is modified before translation.
Introns are removed and exons are spliced together to form mature mRNA.
A 5' cap is added.
A 3' polyA tail is added.
These modifications help stabilize mRNA.
Alternative splicing allows different combinations of exons to be joined.
This means one gene can produce different polypeptides, increasing protein diversity without increasing gene number.
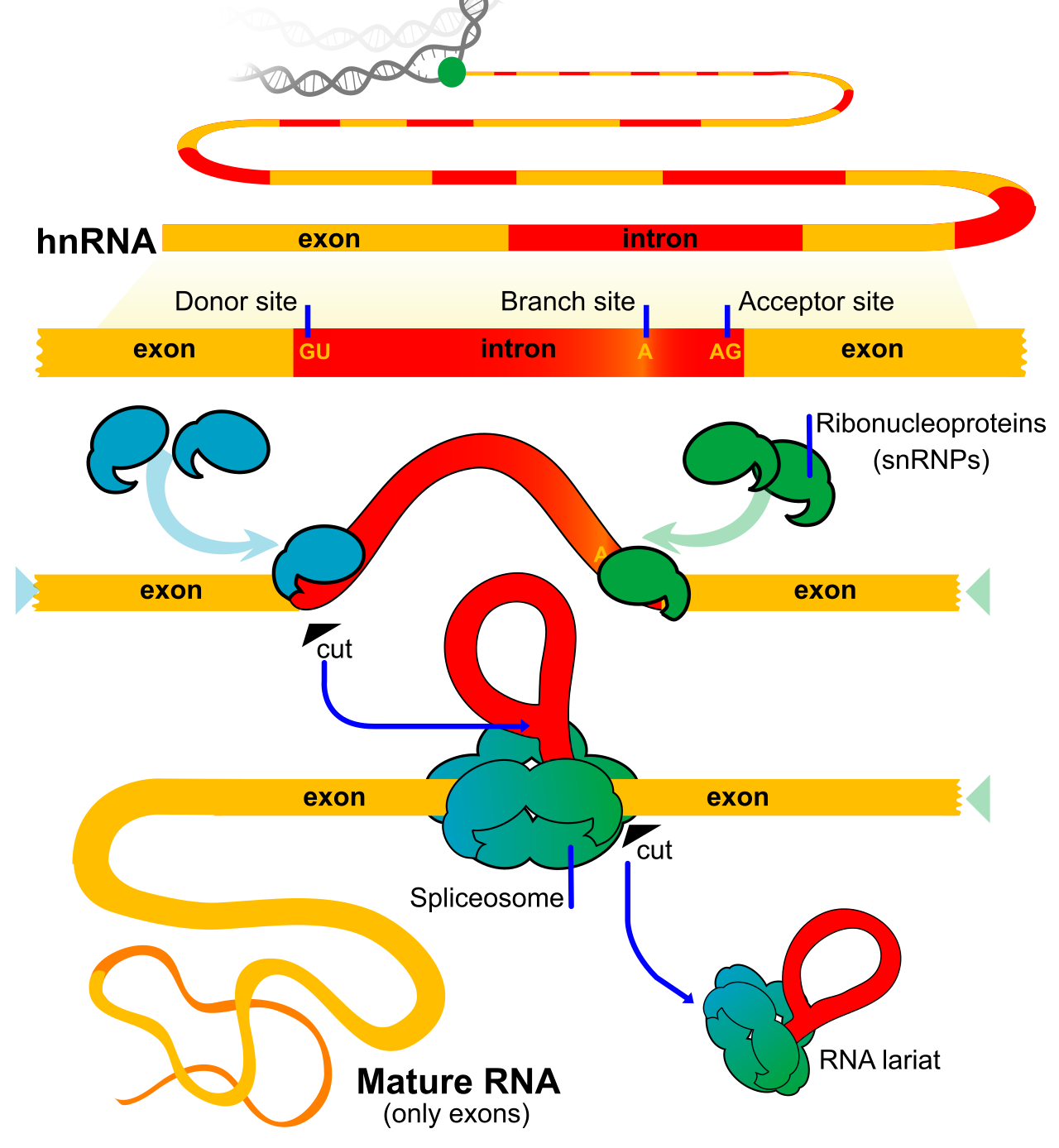
This diagram shows introns being removed and exons being joined to make mature mRNA. It also supports explanations of how alternative splicing can produce different protein variants from one gene. Source
HL only: translation initiation and ribosome details
Translation initiation starts when the small ribosomal subunit attaches to the 5' end of mRNA.
It moves to the start codon.
The initiator tRNA binds first.
Another tRNA binds, then the large ribosomal subunit joins.
The ribosome has three tRNA binding sites: A, P, and E.
During elongation:
A site = incoming aminoacyl-tRNA.
P site = tRNA holding the growing polypeptide.
E site = exit site for discharged tRNA.
Understanding the A, P, and E sites helps explain ribosome movement and peptide bond formation in more detailed HL questions.
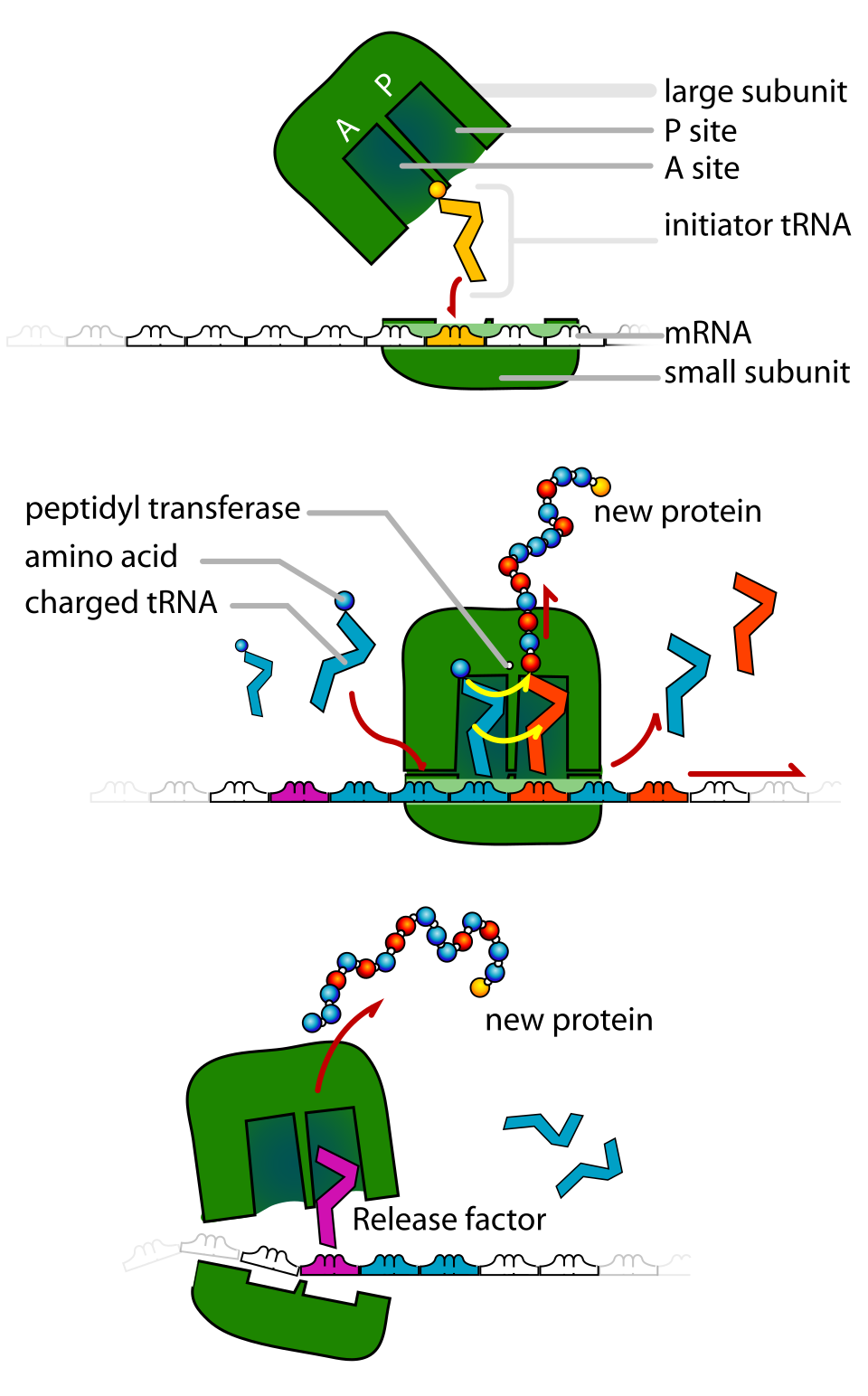
This image clearly shows tRNA molecules interacting with the ribosome and mRNA. It is especially useful for HL students learning the roles of the A, P, and E sites during elongation. Source
HL only: protein processing after translation
Many newly made polypeptides are not yet functional immediately after translation.
They may need modification to become their final functional form.
A required example is insulin:
Preproinsulin is made first.
It is processed to proinsulin.
It is then modified to form insulin.
This shows that the final active protein may be produced only after post-translational modification.
The cell also continually breaks down proteins using proteasomes.
Proteasomes recycle amino acids, which can then be reused in further protein synthesis.
This constant breakdown and resynthesis is necessary to maintain a functional proteome.
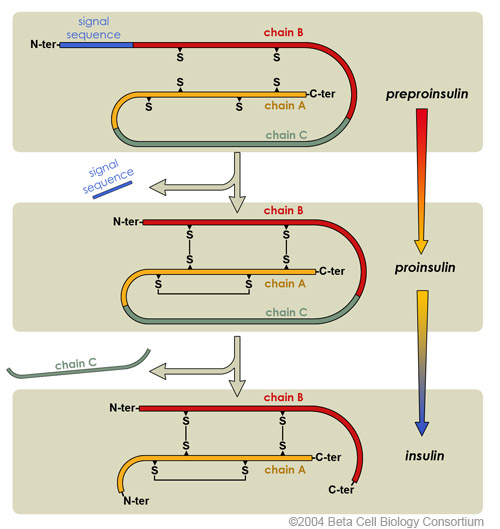
This image shows the conversion of preproinsulin to proinsulin and then to insulin. It is a strong example of post-translational modification, showing that translation alone does not always produce the final active protein. Source
Common exam links and comparisons
Transcription uses DNA as template and produces RNA.
Translation uses mRNA as template and produces a polypeptide.
Complementary base pairing is used in both processes, but the pairings differ slightly because RNA contains uracil (U) instead of thymine (T).
DNA stores the information; mRNA carries the message; tRNA decodes the message; ribosomes assemble the polypeptide.
Mutations matter because they can alter codons and therefore alter amino acid sequence and protein function.
In eukaryotes, extra complexity comes from RNA processing, alternative splicing, and post-translational modification.
Checklist: can you do this?
Explain the difference between transcription and translation, including the role of RNA polymerase, mRNA, tRNA, and ribosomes.
Use a codon table to convert an mRNA sequence into an amino acid sequence.
Identify codons, anticodons, peptide bonds, and the direction 5' → 3' in diagrams or data questions.
Predict how a point mutation could change a polypeptide and therefore alter protein function.
Describe HL processes such as promoter binding, splicing, alternative splicing, translation initiation, and post-translational modification.

Shubhi is a seasoned educational specialist with a sharp focus on IB, A-level, GCSE, AP, and MCAT sciences. With 6+ years of expertise, she excels in advanced curriculum guidance and creating precise educational resources, ensuring expert instruction and deep student comprehension of complex science concepts.
Shubhi is a seasoned educational specialist with a sharp focus on IB, A-level, GCSE, AP, and MCAT sciences. With 6+ years of expertise, she excels in advanced curriculum guidance and creating precise educational resources, ensuring expert instruction and deep student comprehension of complex science concepts.